Research Solution
A customized proteome analysis solution designed to support research,
performing protein analysis on various samples such as blood, tissue, and cells.
- Profiling
: Blood proteome - Profiling
: Tissue/Cell proteome
Deep profiling
- An advanced analytical platform that enables in-depth interpretation of the blood proteome.
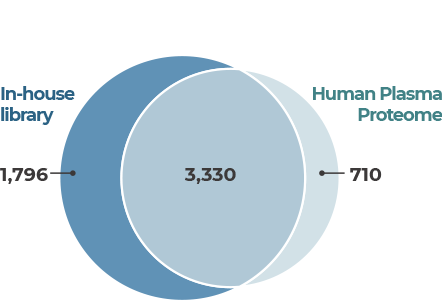
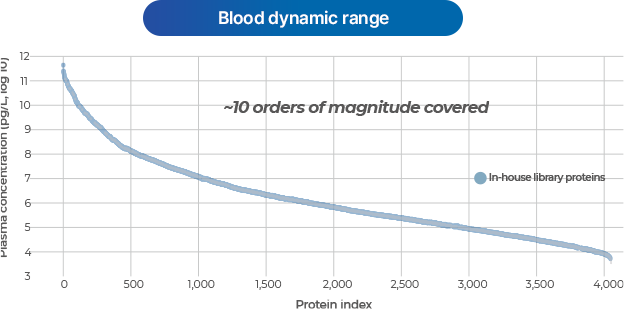
Depletion
- Removes high-abundance proteins in blood to discover even low-abundance proteins in plasma or serum.
-
- 1 Antibody
-
Uses antibodies to remove high-abundance proteins (e.g., Albumin, IgG) for deeper detection.
(Human: 14 proteins depletion, Mouse: 3 proteins depletion)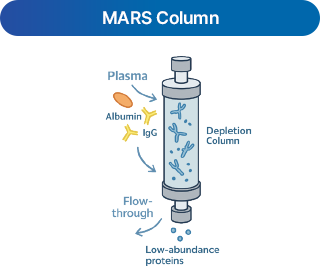
-
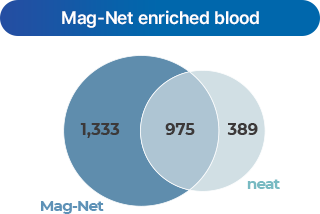
- 2 SAX magnetic bead
-
Automated low-abundance protein analysis using SAX (Strong Anion Exchange) magnetic beads.
Olink Reveal (Proximity Extension Assay)
- Olink Reveal enables discovery of over a thousand proteins from small blood volumes using a PEA-based (Proximity Extension Assay) platform.
- When combined with mass spectrometry-based profiling, it provides more comprehensive proteome information

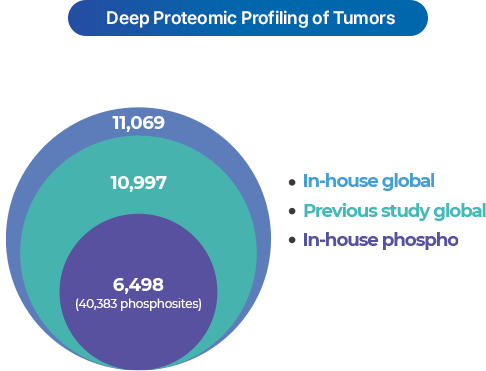
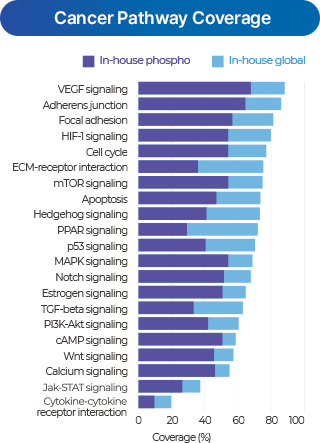
Deep profiling
- Applies automated fractionation technology (COFFER) to evenly separate peptides, ensuring high reproducibility and sensitivity for deep profiling.
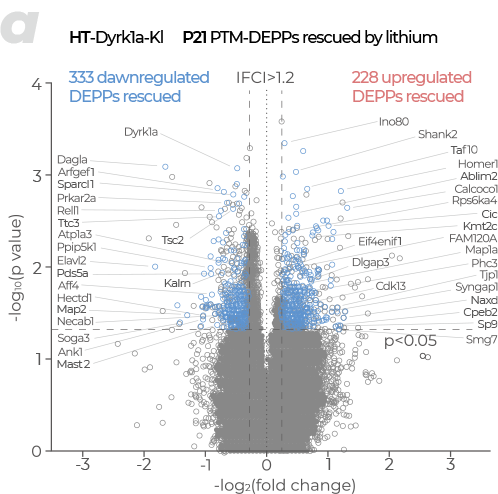
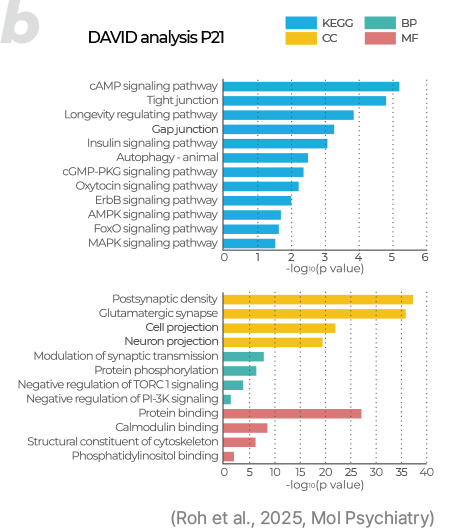
PTM (Post-Translational Modification)
- Provides analysis of diverse PTMs not detectable by RNA analysis, including phosphorylation, acetylation, methylation, ubiquitination, and glycosylation, offering both qualitative and quantitative results.
INTERACTOME
- Performs mass spectrometry on samples from various IP methods (antibody, tag, labeling, etc.) to identify and quantify proteins interacting with a target protein (bait).
- Essential for understanding biological mechanisms such as cell signaling, complex formation, and functional regulation.
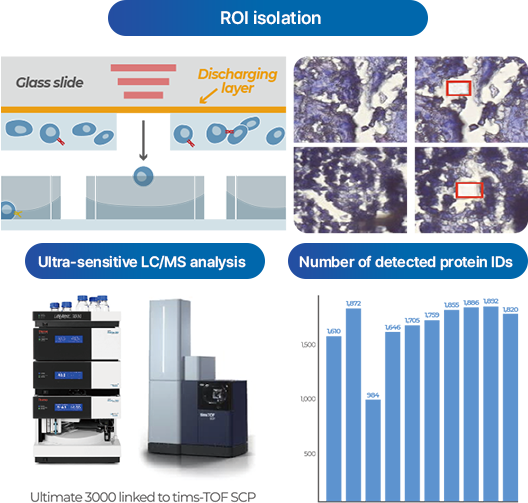
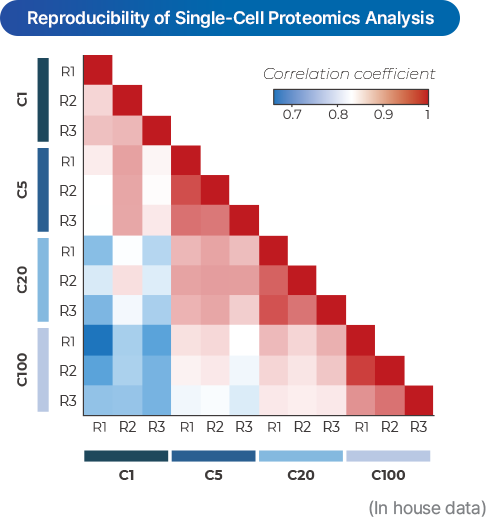
SPATIAL SINGLE-CELL PROTEOMICS
- PASS uniquely applies ultra-sensitive Astral and timsTOF SCP mass spectrometry technologies in Korea to provide single-cell level proteome analysis (>1,000 proteins).
- Applicable to cell lines, in-vitro samples, and clinical tissue samples (FFPE).
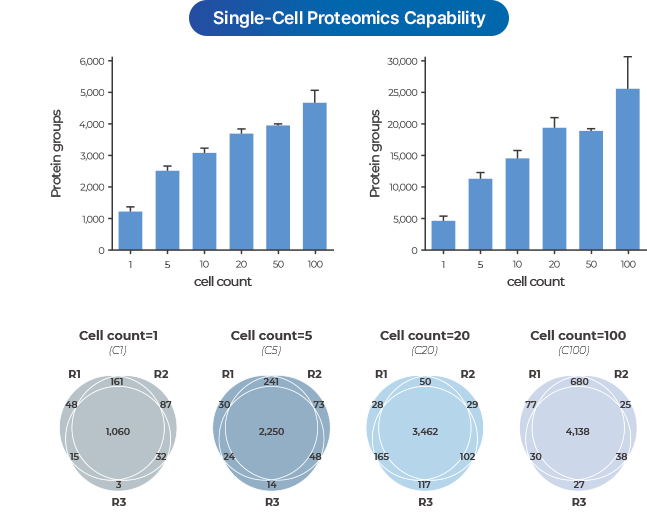
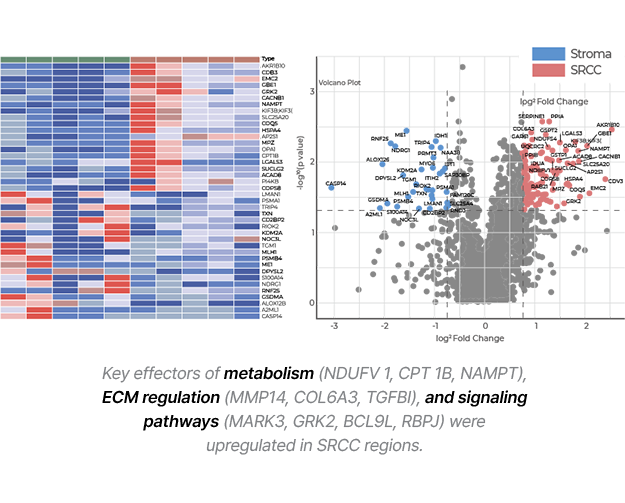
- In gastric cancer FFPE samples, over 1,500 proteins were reliably detected in precisely separated ROIs (Regions of Interest) at the single-cell level, clearly distinguishing protein expression differences between tumor and surrounding tissues.